Disclaimer: This article is more than two years old. Developments in science and computing happen quickly, and more up-to-date resources on this topic may be available.
In This Issue:
- From the Director
- Lab Impact: Preparing hypre for Sierra and GPU-Based Architectures
- Collaborations: Using Data Science to Advance Cancer Treatments with the Norwegian Government
- Advancing the Discipline: IDEALS: Improving Data Exploration and Analysis at Large Scale
- Path to Exascale: LLNL's Roles in the Exascale Computing Project
- Highlights
From the CASC Director
Welcome to the first edition of CASC's new online newsletter. Through this quarterly forum, we will illustrate the broad range of work that CASC does for you: our collaborators, sponsors, staff members, or any computer science and mathematics enthusiast who finds us here.
With 110 scientists working in a dozen major program areas, we cover a lot of ground. Our collaboration with the NNSA Advanced Simulation and Computing (ASC) program goes back to the very beginning of CASC, and our people continue to play central roles in many ASC projects, as well as virtually every major Program within the Laboratory. Meanwhile, 53 members of CASC worked on 39 different LDRD (internally funded Laboratory research) projects just in the first half of Fiscal Year 2017. The depth and the breadth of CASC's collaborations are both signs of success. However they also create a challenge for us: How do we identify our major accomplishments? What unites CASC as an organization?
The CASC newsletter will begin to answer these questions by highlighting in each issue elements of what we see as our core values: Lab Impact through our contributions to the LLNL mission, Collaborations with external partners, Advancing the Discipline through innovative and impactful research, and The Path to Exascale outlining the various roles CASC is playing in helping the US and DOE overcome the challenges of next-generation HPC architectures. We can't hope to cover the whole range of our work in a few issues or even a few years, but we hope that each issue will give you a sense of our values, our priorities, our accomplishments, and our people.
If you've come across this newsletter through your favorite social media outlet, please give it like, share, or retweet to help us get the word out.
Contact: John May
Lab Impact | Preparing hypre for Sierra and GPU-Based Architectures
The hypre library has been one of CASC's signature projects for nearly 20 years. It focuses on the parallel solution of linear systems at extreme scale and features multigrid methods for scalability. One of the unique aspects of hypre is that it provides multigrid solvers for both unstructured and structured problems by way of its conceptual linear system interfaces. The algebraic multigrid solver BoomerAMG is the workhorse solver in hypre developed mainly for unstructured problems. For structured problems, the semi-coarsening multigrid methods PFMG and SMG are available.
The hypre team has continually adapted the library to new architectures, programming models, and user need. Today, the Lab’s ASC and institutional code teams are preparing for the deployment of Sierra. In support of these codes, the hypre team is focusing on the development of GPU-enabled solvers as part of the iCOE Libraries and Tools project led by CASC's Ulrike Yang. Another important aspect of the team’s work is the development of new math algorithms that improve convergence, increase concurrency, and reduce communication when implemented in parallel. To date, hypre has relied on a hybrid MPI+OpenMP parallel computing model. The GPU work experiments with several approaches.
In the structured component of the library, solver kernels already utilized a loop abstraction called BoxLoop. The use of this abstraction made it somewhat straightforward to implement a variety of approaches for programming GPUs, including CUDA, OpenMP 4.X, RAJA, and Kokkos. This approach primarily involved writing different implementations of the BoxLoop macros themselves and was not overly intrusive on the existing code base. Speedups of up to 2x for SMG and 8x for PFMG were observed on Ray versus the CPU-only code. The larger speedups occurred for larger problem sizes. The most important aspect for achieving good performance was careful memory management by minimizing data motion between the CPU and GPU.
For the unstructured component of the library, the algebraic multigrid solver BoomerAMG has been the focus. Every solver in hypre has a setup and a solve phase, but the setup phase in BoomerAMG is somewhat complex. For this reason, although considerable effort has already been put into the setup phase, the focus is primarily on the solve phase. The main kernel in the solve phase is matrix-vector multiplication, so efforts were made to port that kernel to GPUs through a variety of routes, including Cuda, OpenMP 4.X, RAJA, and routines from Nvidia’s CuSparse library. Speedups of up to 9x were observed versus the CPU-only code for the solve phase on larger problems. As with the structured solvers, careful memory management proved to be critical to achieving good performance.
The hypre team is about to release the first version of the library with support for GPUs and will then coordinate with the ASC code groups to run tests in the context of representative simulations. These tests will help guide implementation improvements. The team will also work with the SUNDIALS and MFEM projects to test use of hypre with these libraries and investigate ways to deal with upcoming issues.
Contact: Rob Falgout
Collaborations | Using Data Science to Advance Cancer Treatments with the Norwegian Government
Delivering care tailored to the needs of the individual, rather than population averages, has the potential to transform the delivery of healthcare. The convergence of high performance computing, big data and life science is enhancing the development of personalized medicine.
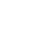
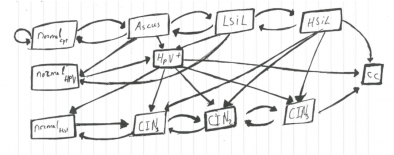
Our team is partnering with the Cancer Registry of Norway to advance “precision medicine” by developing individually tailored prevention and treatment strategies. The partnership’s goal is to improve the outcome of women affected by cervical and breast cancers.
Norway provides equal access to health care to all citizens and maintains registries of health histories. As a result, Norway brings an invaluable resource to the collaboration, a national database for some 1.8 million ‘unique’ Norwegian women’s cervical cancer screening results covering 25 years (1991-2015). Aggressive screening programs against cancer, begun in 1995, are a key element of the Norwegian government’s cancer control effort.
Our initial efforts are focused on improving risk assessment and the resulting screening recommendations for individual women. We are working to personalize screenings by combining pattern recognition, machine learning and time-series statistics to analyze the data. Access to 10.7 million individual records for 1.8 million patients over 25 years provides the opportunity to test and validate our models against historical data. The team is developing a flexible, extendable model that incorporates new data such as other biomolecular markers, genetics and lifestyle factors to individualize risk assessment. Reliable assessment would allow women with increased risk of cervical cancer to receive more frequent screenings and those with lower risk to receive fewer, improving health outcomes and making screening programs more cost effective. Medical researchers, who are part of our team, note that increasing the time between screenings for low-risk women reduces the harm associated with false-positive tests such as anxiety and unnecessary medical costs.
We addressed the absence of privacy and security protocols between the two countries for exchanging medical data. We were given access to the data in March of 2016, and early work shows promise. We model the data using Hidden Markov Model (HMM) and our first prototype was developed using R. We are currently implementing a Python version to address the scalability of the model. Currently the partnership’s focus is on cervical cancer, and the plan is to later include breast cancer as well.
Contact: Ghaleb Abdulla
Advancing the Discipline | IDEALS: Improving Data Exploration and Analysis at Large Scale
The quantitative analysis of small to moderate-sized datasets using statistical and machine learning techniques, while far from trivial, is often tractable. The small size of the data allows us to examine them, understand their properties, devise a solution approach, evaluate the results, and refine the analysis. This gives us confidence in the results. In contrast, when we first encounter a massive dataset, the sheer size of the data can be overwhelming, making it challenging to fully explore the data and evaluate an analysis approach. Consequently, we could select less-than-optimal algorithms for the analysis, increasing the risk of drawing incorrect conclusions from the data. To address these concerns, the IDEALS project was recently funded under the Scientific Data Management, Analysis, and Visualization at Extreme Scale (SDMA&V) program in ASCR/DOE.
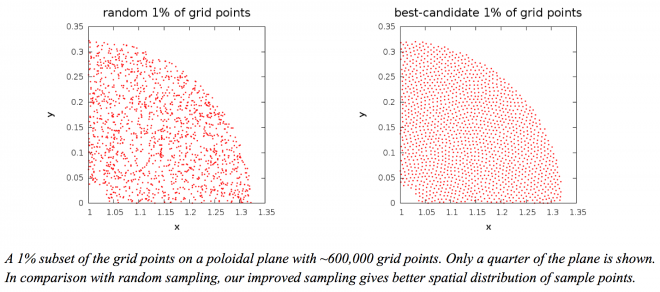
The objective of IDEALS (Improving Data Exploration and Analysis at Large Scale) is to enable scientists to gain confidence in the analysis of large-scale data sets. Our initial focus has been on simple ways to explore large data sets. An obvious solution is to work with a smaller sample of the data. This smaller subset must be selected randomly to reduce any bias. However, it is well known that a random sampling leads to regions that are over- or under-sampled. We are investigating alternate techniques that generate more uniformly distributed samples. We are particularly interested in techniques that require little memory, are flexible so they can be applied to data in different data structures, and are efficient so they can provide near-real-time response on large datasets. The figure below shows a 1% sample of the grid points in a quarter of a poloidal plane from a fusion simulation. We devised a one-pass version of an improved sampling algorithm that gives a better spatial distribution of the selected subset of grid points. We are currently investigating how we can use this subset to explore the data further.
Contact: Chandrika Kamath
Path to Exascale | LLNL's Roles in the Exascale Computing Project
Since the first petascale (10^15 operations/second) HPC systems arrived at Los Alamos in 2008, the DOE has been anticipating the imminent challenges facing the HPC community in achieving the next iconic 1000x factor of performance improvement known as exascale. With the official launch of the Exascale Computing Project (ECP) in 2016, LLNL and CASC are at the forefront of activities aimed at making that goal a reality. CASC is primarily involved in the Software Technologies focus area in the ECP, which aims to develop a production-ready and comprehensive software stack in support of ECP goals. CASC personnel are also embedded in LLNL Programs who are developing applications ready to run on exascale systems, as well as Livermore Computing who is preparing to house one of the first U.S.-based exascale platforms.
The ECP software stack is organized around eight topical areas spanning the software infrastructure needs of the HPC community, and those projects will be the focus of future articles in this newsletter. The projects for which LLNL is the lead laboratory in each area include:
Programming Models and Runtimes
- Participating in MPI and OpenMP standardization efforts
- Integrated Software Components for Managing Computation and Memory Interplay at Exascale
- Runtime System for Application-Level Power Steering on Exascale Systems
- Using ROSE to help transition ASC application source code to exascale-ready
Tools
- Hardening and productization of Performance Tools
- Next-Generation Computing Environment software suite
- Exascale Code Generation Toolkit
Math Libraries and Frameworks
Data Management and Workflows
- Data Management and Workflow/Volume Rendering
- UNIFYCR: A Checkpoint/Restart File System for Distributed Burst Buffers
- ZFP: Compressed Floating-Point Arrays
Data Analytics and Visualization
- Integration of Volume Rendering to an Exascale Application
System Software
- Flux Resource Management Framework
- HPC Developer's Environment
Co-Design and Integration
- Advanced Architecture Portability Specialists
The ECP highlights the importance of collaboration in what is certainly the most ambitious HPC effort ever to be undertaken in the DOE, and most, if not all, of the projects above involve partners from other DOE laboratories and/or universities. Likewise, LLNL is a partner in a large number of other ECP projects led by other institutions.
CASC is also leading the deployment of an ECP Proxy App Suite under the Application Development focus area of ECP, which will collect and curate a set of small representative applications for use in co-design with hardware vendors and software stack developers.
Future editions of this newsletter will highlight many of the above CASC projects as they progress along the path to exascale. Stay tuned to these pages for more information.
Contact: Rob Neely
CASC Highlights
New Hires (since January 1, 2017)
- Stephan Gelever (Portland State University)
- Abishek Jain (Nanyang Technical University)
- Sookyung Kim (Georgia Institute of Technology)
- Timothy La Fond (Purdue University)
- Shusen Liu (University of Utah)
- Naoya Maruyama (RIKEN Advanced Institute for Computer Science)
- Lee Ricketson (Courant Institute and UCLA)
Awards
- Carol Woodward was named a Fellow of the Society for Industrial and Applied Mathematics (SIAM) "for the development and application of numerical algorithms and software for large-scale simulations of complex physical phenomena."
- Ghaleb Abdulla was given a Secretary's Appreciation Award by former Energy Secretary Ernest Moniz "for responding to the Vice President's Cancer Moonshot to make a decade of progress in five years in the war on cancer through innovative partnerships and approaches with government agencies, industry, and foreign countries."
Looking back...
20 years ago CASC celebrated its first anniversary with a picnic outside its home building. LLNL's Newsline noted CASC's rapid growth over the first year, from 15 scientists to more than 30. The article lists CASC's core competencies as including HPC, computational physics, numerical mathematics, algorithm development, scientific data management, and visualization. That list would be nearly identical today.
Help us out!
This newsletter is looking for a permanent name. If you have and idea for a catchy name, please send them along to our admins before May 31st. If your idea is selected, we'll send you a $10 Starbucks gift card!
Or, if you're aware of a CASC project that you'd like to see highlighted in our next issue (coming out this summer), let us know.