Browse this site's news, projects, and people highlights via any of the topics in the dropdown list or below each content description.
Ardra
Researchers are testing and enhancing a neutral particle transport code and its algorithm to ensure that they successfully scale to larger and more complex computing systems.
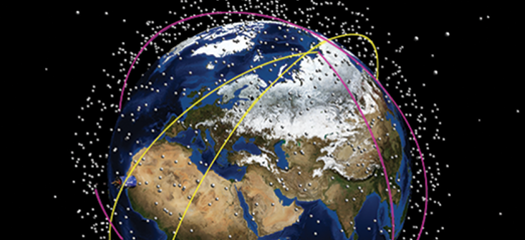
TESSA
Testbed Environment for Space Situational Awareness software helps to track satellites and space debris and prevent collisions.

Serpentine Wave Propagation
These methods for solving hyperbolic wave propagation problems allow for complex geometries, realistic boundary and interface conditions, and arbitrary heterogeneous material properties.

LLNL, DOD, NNSA dedicate Rapid Response Laboratory and supercomputing system to accelerate biodefense
The collaboration has enabled expanding systems of the same architecture as LLNL’s upcoming exascale supercomputer, El Capitan, featuring AMD’s cutting-edge MI300A processors.

Understanding the changing climate
Ensuring researchers and policymakers can predict and prepare for the long-term effects of a changing climate is a central, motivating question for the Energy Exascale Earth System Model (E3SM) project.

Department of Energy announces FASST initiative
The proposed Frontiers in Artificial Intelligence for Science, Security and Technology (FASST) initiative will advance national security; attract and build a talented workforce; harness AI for scientific discovery; address energy challenges; develop technical expertise necessary for AI governance.
